AINeutralarXiv – CS AI · 3d ago7/10
🧠BioArc introduces a neural architecture search framework that systematically discovers optimal model architectures for biological foundation models, moving beyond generic adaptation of NLP and computer vision models. The research identifies design principles and proposes methods to predict architectures for new biological tasks, providing foundational methodology for next-generation biology-focused AI systems.
AIBullisharXiv – CS AI · 5d ago7/10
🧠Researchers introduce AIMS-Fold, a guided-diffusion framework that integrates structural proteomics data (XL-MS and HDX-MS measurements) with protein structure prediction models to improve accuracy in predicting protein complex conformations. The approach outperforms unguided computational models on challenging induced proximity drug targets, advancing structure-based drug design capabilities.
AIBullisharXiv – CS AI · May 127/10
🧠Researchers introduce Yeti, a compact protein structure tokenizer that converts protein structures into discrete tokens for multimodal AI models. The approach achieves superior codebook utilization and token diversity while maintaining competitive reconstruction accuracy with 10x fewer parameters than existing solutions, enabling efficient joint generation of protein sequences and structures.
AIBullishGoogle DeepMind Blog · Nov 257/102
🧠AlphaFold has significantly accelerated scientific research and biological discovery over the past five years. The AI system has enabled breakthroughs in protein structure prediction, fueling innovation across the global scientific community.
AIBullishNVIDIA AI Blog · Feb 197/102
🧠NVIDIA has made Evo 2, the largest publicly available AI foundation model for genomic data, accessible through its BioNeMo platform. The model was developed in collaboration with Arc Institute and can understand genetic code across all domains of life, built on NVIDIA's DGX Cloud platform.
AIBullisharXiv – CS AI · 4d ago6/10
🧠Researchers introduce ProtLiD², a discrete diffusion model that co-designs protein sequences and structures while conditioning on ligand information, achieving significant improvements in fold confidence and ligand-binding accuracy compared to existing methods. The model demonstrates practical advantages in both whole-protein and active-site pocket design tasks.
🏢 Meta
AINeutralarXiv – CS AI · 5d ago6/10
🧠Researchers introduce SILO, a self-improvement imitation framework for protein design that optimizes protein sequences under limited evaluation budgets. The method combines hierarchical editing, stochastic beam search, and active learning to outperform existing reinforcement learning and generative approaches across multiple protein fitness landscapes.
AINeutralarXiv – CS AI · 5d ago6/10
🧠Researchers introduce TriProRep, a protein representation learning method that jointly models amino acid identity, backbone geometry, and full-atom geometry to improve protein structure prediction. The new approach outperforms sequence-only and prior structure-aware models across multiple benchmarks including homodimer co-folding and monomer structure prediction tasks.
AIBullishMIT News – AI · May 206/10
🧠Connor Coley is advancing machine learning applications in chemistry to accelerate drug discovery and compound design. This work represents a convergence of AI with pharmaceutical research, enabling computational models to understand and predict chemical behavior more effectively than traditional methods.
AINeutralGoogle DeepMind Blog · May 166/10
🧠Filippo Menolascina leverages Co-Scientist AI to accelerate the discovery of liver disease mechanisms and identify new treatment options. The research aims to explain why certain existing drugs are effective only for specific patient populations, potentially enabling more personalized therapeutic approaches.
AINeutralarXiv – CS AI · May 126/10
🧠Researchers propose L3-PPI, a biologically-informed machine learning approach for predicting protein-protein interactions by leveraging the L3 rule—the principle that multiple length-3 paths between proteins indicate interaction likelihood. The method integrates a lightweight graph prompt learning module into existing PPI predictors as a plug-and-play component, demonstrating superior performance over conventional approaches that rely on generic aggregation methods.
AIBullisharXiv – CS AI · May 126/10
🧠Researchers introduce improved methods for Gene Regulatory Network (GRN) inference using single-cell foundation models, proposing Virtual Value Perturbation and Gradient Trajectory techniques to better extract regulatory knowledge. The work establishes a new benchmark for evaluating GRN predictions across unseen genes and datasets, demonstrating significant performance improvements over existing approaches.
AIBullisharXiv – CS AI · May 116/10
🧠Researchers have developed a novel discrete diffusion model that improves computational antibody design by using germline sequences as an anchor point rather than masked tokens, reducing memorization of genetic patterns and enabling better conditional generation of antibodies with specific therapeutic properties like improved binding affinity.
AIBullisharXiv – CS AI · May 116/10
🧠Researchers demonstrate that ProteinJEPA, a latent-space prediction technique, can complement traditional masked language modeling (MLM) in protein language models, achieving better downstream task performance when combined strategically. The optimal approach—masked-position MLM+JEPA—wins 10 out of 16 evaluation tasks against MLM-only baselines while maintaining computational efficiency.
AINeutralarXiv – CS AI · May 116/10
🧠Researchers introduce BeeVe, an unsupervised machine learning framework that discovers acoustic patterns in honey bee hive sounds without labels or predefined categories. The system successfully identifies distinct behavioral states linked to hive health conditions, demonstrating that AI can extract meaningful biological structure from non-vocal animal signals.
AIBullisharXiv – CS AI · Apr 106/10
🧠Researchers introduce MAT-Cell, a neuro-symbolic AI framework that combines large language models with biological constraints to improve single-cell annotation accuracy. The system uses multi-agent reasoning and verification processes to overcome limitations in both supervised learning and LLM-based approaches, demonstrating superior performance on cross-species benchmarks.
AIBullishMarkTechPost · Apr 56/10
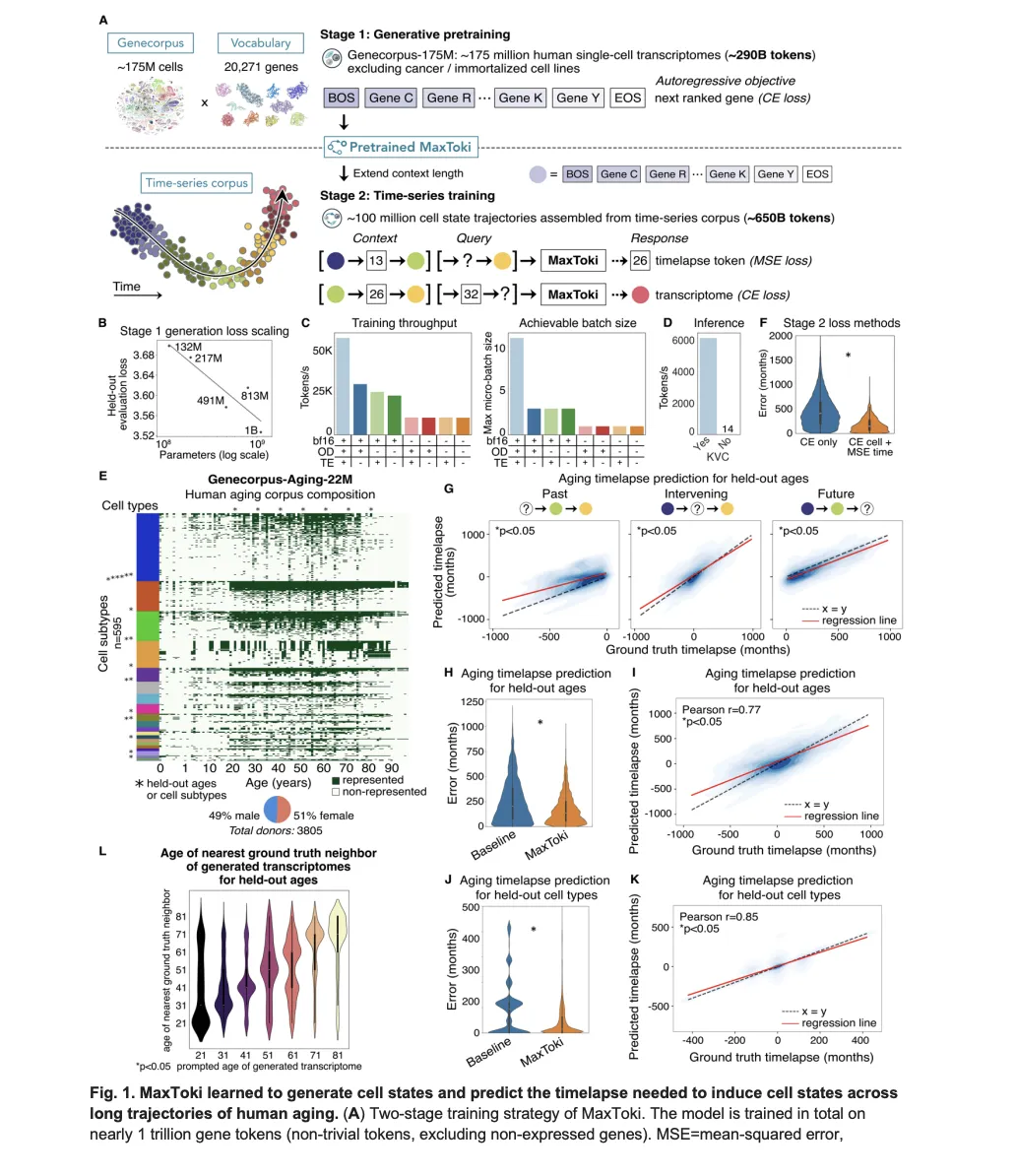
🧠MaxToki is a new AI foundation model that can predict cellular aging patterns and trajectories, addressing a key limitation in existing biological models that only analyze cells as static snapshots. The technology represents a significant advancement in computational biology by incorporating temporal dynamics into cellular analysis.
AIBullisharXiv – CS AI · Mar 36/107
🧠Researchers developed a pharmacology knowledge graph for drug repurposing and found that removing chemical structure representations improved performance while dramatically reducing computational requirements. The study showed that drug behavior can be accurately predicted using only target protein information and network topology, with larger datasets proving more valuable than complex models.
AIBullisharXiv – CS AI · Mar 36/104
🧠Researchers propose a new iterative distillation framework for fine-tuning diffusion models in biomolecular design that optimizes for specific reward functions. The method addresses stability and efficiency issues in existing reinforcement learning approaches by using off-policy data collection and KL divergence minimization for improved training stability.
AINeutralarXiv – CS AI · Mar 175/10
🧠Researchers conducted the first empirical study analyzing how natural scientists reuse pre-trained deep learning models across 17,511 peer-reviewed papers from 2000-2025. The study found that biochemistry and molecular biology lead in model reuse, with adaptation being the most common reuse pattern, primarily impacting the testing phase of scientific research.
AINeutralarXiv – CS AI · Mar 175/10
🧠Researchers developed a comprehensive benchmarking system to evaluate AI agent performance in single-cell omics analysis, testing 50 real-world tasks across multiple frameworks. The study found that Grok3-beta achieved state-of-the-art performance, while multi-agent frameworks significantly outperformed single-agent approaches through specialized role division.
🧠 Grok
AINeutralarXiv – CS AI · Feb 274/106
🧠Researchers developed MEDNA-DFM, a dual-view deep learning model that predicts DNA methylation patterns while providing biological explanations. The model achieves high accuracy across species and includes explainable AI features that reveal conserved genetic motifs and cooperative sequence-structure relationships.
AIBullishMIT News – AI · Feb 44/105
🧠Professor James Collins discusses how collaboration has been central to his research combining computational predictions with experimental platforms to accelerate therapeutic drug discovery and design using AI technologies.
AINeutralMIT News – AI · Dec 154/105
🧠MIT Assistant Professor Yunha Hwang uses computational methods to study microbial genomes and understand biological language. Her appointment demonstrates MIT's focus on combining genetics research with artificial intelligence applications.
AIBullishHugging Face Blog · Jul 35/105
🧠Intel has developed optimizations to accelerate the ProtST protein language model on their Gaudi 2 AI accelerator hardware. This advancement demonstrates Intel's commitment to supporting specialized AI workloads in computational biology and scientific research applications.